Harden Hardin Harding yDNA Project
TIME to MOST RECENT COMMON ANCESTOR
(TMRCA)
The initial objective for participation in a Surname
Project is to use the evaluation of the genetic markers on the Y-chromosome to
establish that two people share a common ancestor. Once this initial goal
has been established, the next goal is then to use the genetic markers to
establish the Time to the Most Recent Common Ancestor (TMRCA). In other
words, how many generations the two individuals are from a common ancestor.
The basic premise is fairly simple: Individuals who match a higher number of markers are more closely related. Imagine the Y-chromosome as a clock that ticks very slowly; i.e., one tick of the genetic clock equals one mutation. Thus, a Y-chromosome is a molecular clock that ticks randomly within a specified rate. This paradoxical sounding phrase means that a clock running longer has a higher probability of having more ticks than a clock that has been running for a shorter time. The more time, the more ticks and the longer time it is back to the MRCA.
Therefore, the TMRCA is based on the observed number of
mutations by which two Y-chromosomes differ. Since mutations occur at
random, the calculation of the TMRCA is not an exact science; e.g., 7
generations, but rather a probability distribution, a function that gives the
probability that the TMRCA is a certain number of generations or less;
e.g., a 50% probability that the TMRCA is 16 generations or less. The
graphs provided below depict this function for various number of markers
tested. As more markers are tested, the distribution becomes tighter and
tighter and the calculations for TMRCA have higher precision.
There are two fundamental assumptions we need to make to
deal with to translate an observed number of mutational differences into a
probability distribution for the TMRCA:
![]() We
must count the true number of mutations, and
We
must count the true number of mutations, and
![]() We
must be able to determine the rate of the clock; i.e., assumptions about
the mutation rate.
We
must be able to determine the rate of the clock; i.e., assumptions about
the mutation rate.
If we simply count the number of markers at which two
individuals disagree as the number of mutations, we may run into some
problems. First, some of the markers can differ by one-step, or by
two-steps, or by even more steps. Should we count a two-step difference
as one mutation or two or more mutations? Likewise, even if two markers
appear to be identical for two individuals (normally scored as no mutation),
there is always a small probability that each individual has experienced a
mutation to the same marker since the MRCA and hence the true mutant count for
this particular marker would be two, not zero.
The genetic scientists use two approaches to determine
the number of true mutations.
The Infinite Alleles Model (IAM) is a fancy
population-geneticist term for ‘what you see is what you get’ - the assumption
is that the observed number of mutations equals the true number of mutations.
On the other extreme is the Stepwise Mutational Model (SMM), which corrects for
so-called multiple hits - mutants we might have missed. When the fraction
of matches is very high, both methods provide essentially the same probability
curve. They only differ significantly as individuals become increasingly
dissimilar. For genealogical purposes, we can use the IAM without being
too concerned.
The second issue is setting the clock. This is just a function of the mutation rate. We already know that mutation rates differ for different markers, and markers with higher mutation rates provide faster clocks. Faster clocks are a good thing for us, in that they permit more precision in establishing the TMRCA. We make the initial assumption that the mutation rate is the same for each marker, something that will be adjusted as new data becomes available (note: FTDNA and the University of Arizona geneticists are currently evaluating mutation rates for individual markers). The TMRCA is calculated using two different mutation rates - the standard average over many previous Y- chromosome studies of around 0.002 (1/500) per generation; in other words on the average there is one mutation for an individual marker every 500 generations (that’s approximately 12,000 human years) and a faster rate that is consistent with at least some of the data currently available.
TMRCA GRAPHS
This first TMRCA graph depicts the number of generations (based on a 50% probability) to the common ancestor for two men tested who match all Y-chromosome markers; e.g., 25 for 25. One can readily see from this graph that the more markers that are tested, the fewer number of generations it is to the common ancestor for the two men who have identical Y-DNA profiles. The TMRCA will no doubt be shown to be slightly less than shown by this graph when more data is available concerning mutation rates for individual markers. The TMRCA would be higher for matches involving two men who don't have a perfect match; e.g. 24 for 25.
TMRCA
GRAPH 1
GENERATIONS
to TMRCA vs. NUMBER of MARKERS TESTED (50%
Probability)
|
|
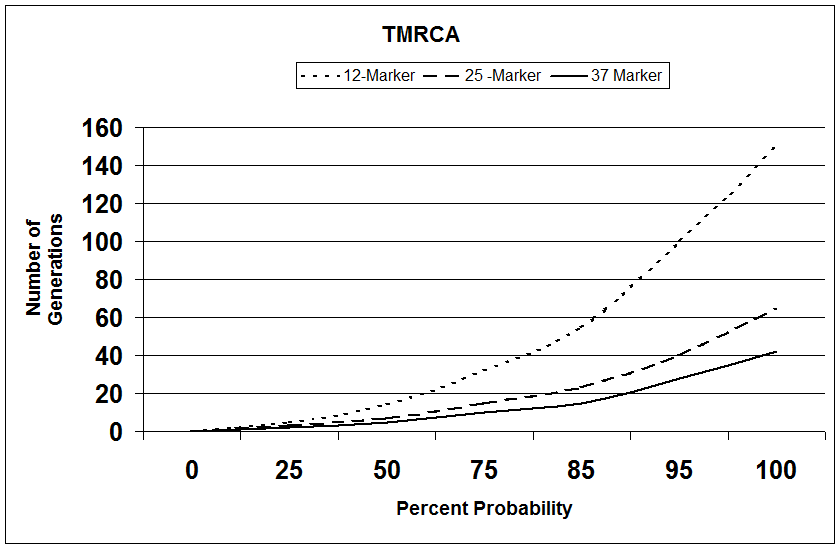
This second TMRCA graph depicts the confidence level for the TMRCA to the common ancestor for two men tested who match all Y-chromosome markers; e.g., 12 for 12, 25 for 25 and 37 for 37. One can readily see from this graph that the more markers that are tested, the higher confidence level for the calculation for the number of generations to the common ancestor for the two men who have identical Y-DNA profiles. The confidence level will no doubt be shown to be slightly higher than shown by this graph when more data is available concerning mutation rates for individual markers. The confidence level would be lower for matches involving two men who don't have a perfect match; e.g. 24 for 25
TMRCA GRAPH 2
CONFIDENCE LEVEL for TMRCA vs.
NUMBER of MARKERS TESTED